HIGH-THROUGHPUT DEEP MUTATIONAL SCANNING (DMS) FOR PREDICTING SARS-COV-2 IMMUNE ESCAPE MUTATIONS
2023-01-18
A world-leading platform for SARS-CoV-2 antibody/ vaccine studies was set up, allowing fast measurement of the immune pressure induced by infection and vaccination.
Capability of quickly assessing the efficacy of established antibody drugs and population immunity against newly emgered variants.
Escape mutation profiles of over 6,000 SARS-CoV-2 antibodies were obtained, enabling us to predict which mutations and variants are the most likely to circulate.
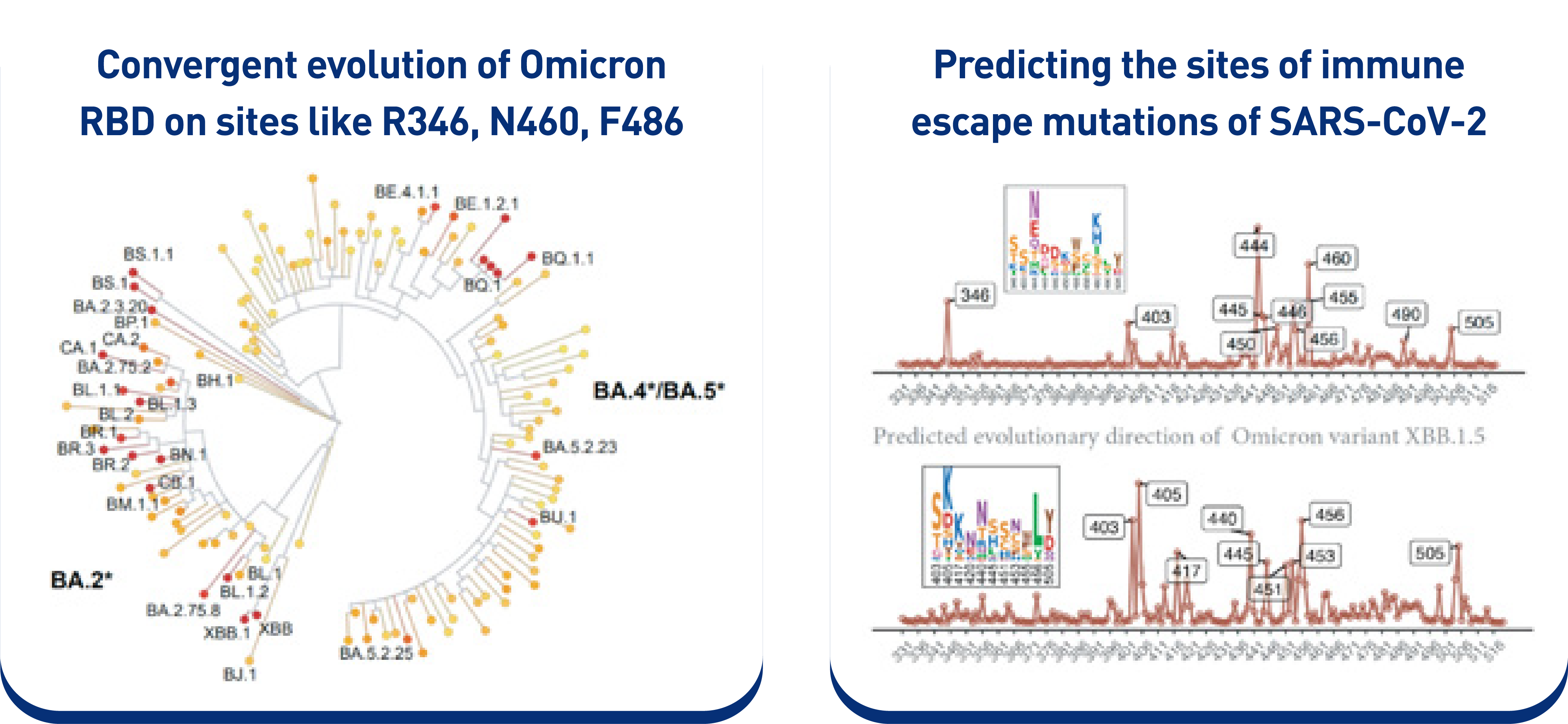
Reveal and rationalize the phenomenon of“convergent evolution” of SARS-CoV-2 Omicron receptor-binding domain (RBD).
MACS-based high-throughput yeast display mutation screening
High-throughput escape-mutations profiling and epitope classification of Nabs
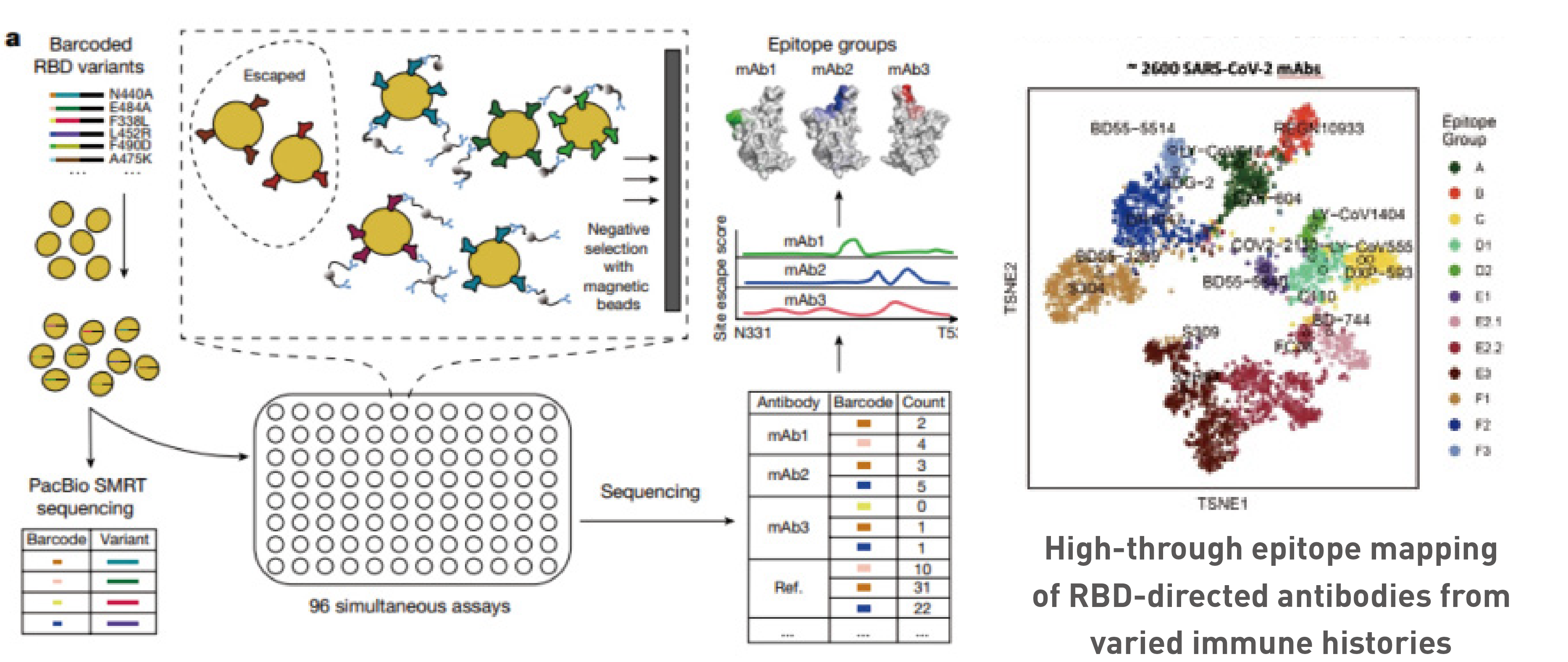
Deciphering humoral immune responses to SARS-CoV-2
Omicron breakthrough infection is limited by immune imprinting

Revealing the humoral immune escape mechanisms of Omicron subvariants: Omicron BA.1 could render most NAb drugs ineffective and cause a substantial reduction in vaccine-induced neutralization titers.
Prospectively alerted the global prevalence of BA.4/5 and BA.2.75: BA.4/BA.5 can escape antibodies induced by BA.1 infection and vaccination; BA.2.75 and BA.5 subvariants can evade the humoral immunity elicited by BA.5 breakthrough infection.
Prospectively alerted the emergence and prevalence of convergent mutants such as XBB.
Predicting the evolution of XBB sublineages: XBB* subvariants carrying 486P mutation (such as XBB.1.5 and XBB.1.9.2) would acquire additional immune-escape mutations due to enhanced hACE2-binding affinity.