GENOME-WIDE FOOTPRINTING OF TRANSCRIPTION FACTORS (TF) WITH DEAMINASE (FOODIE) AND PREDICTION OF ANY TF’S BINDING AFFINITY ON DNA
2026-04-29
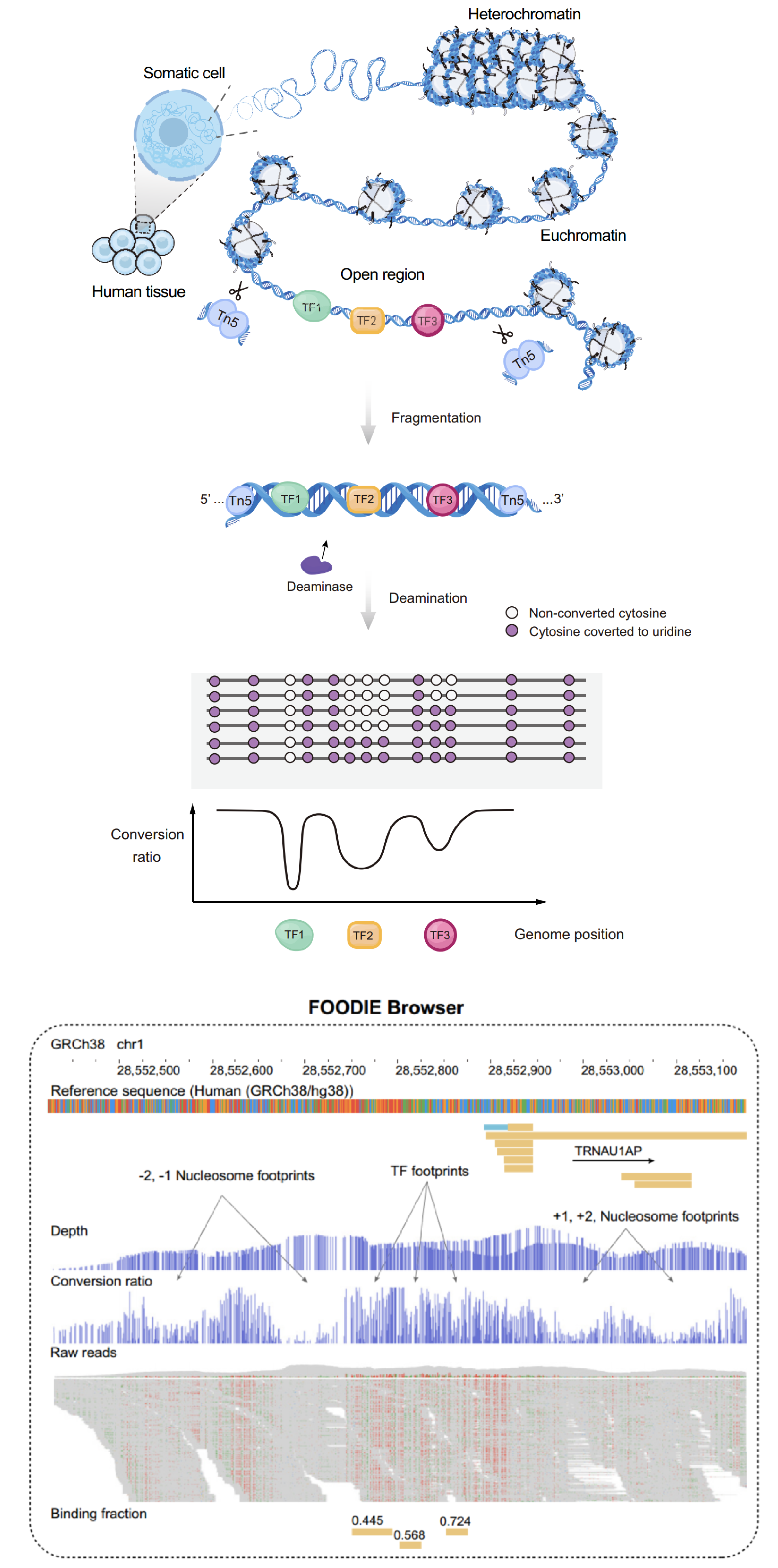
FOODIE is a transformative approach for genome-wide footprinting of transcription factors with deaminase
Transcription factors (TFs) control gene expression by binding DNA at genome’s regulatory regions with sequence-specific affinities. Accurate genome-wide measurements of binding affinities are crucial for understanding regulatory networks but remain experimentally challenging. CPL has developed an innovative technology, FOODIE (FOOtprinting with DEamInasE), a double-stranded DNA deaminase–based transcription factor footprinting method to determine transcription factor (TF)–DNA binding footprints across the entire genome at nearly single-nucleotide resolution. A web-browser is created for visualization of TF footprints along the human genome for different cell lines and tissues.
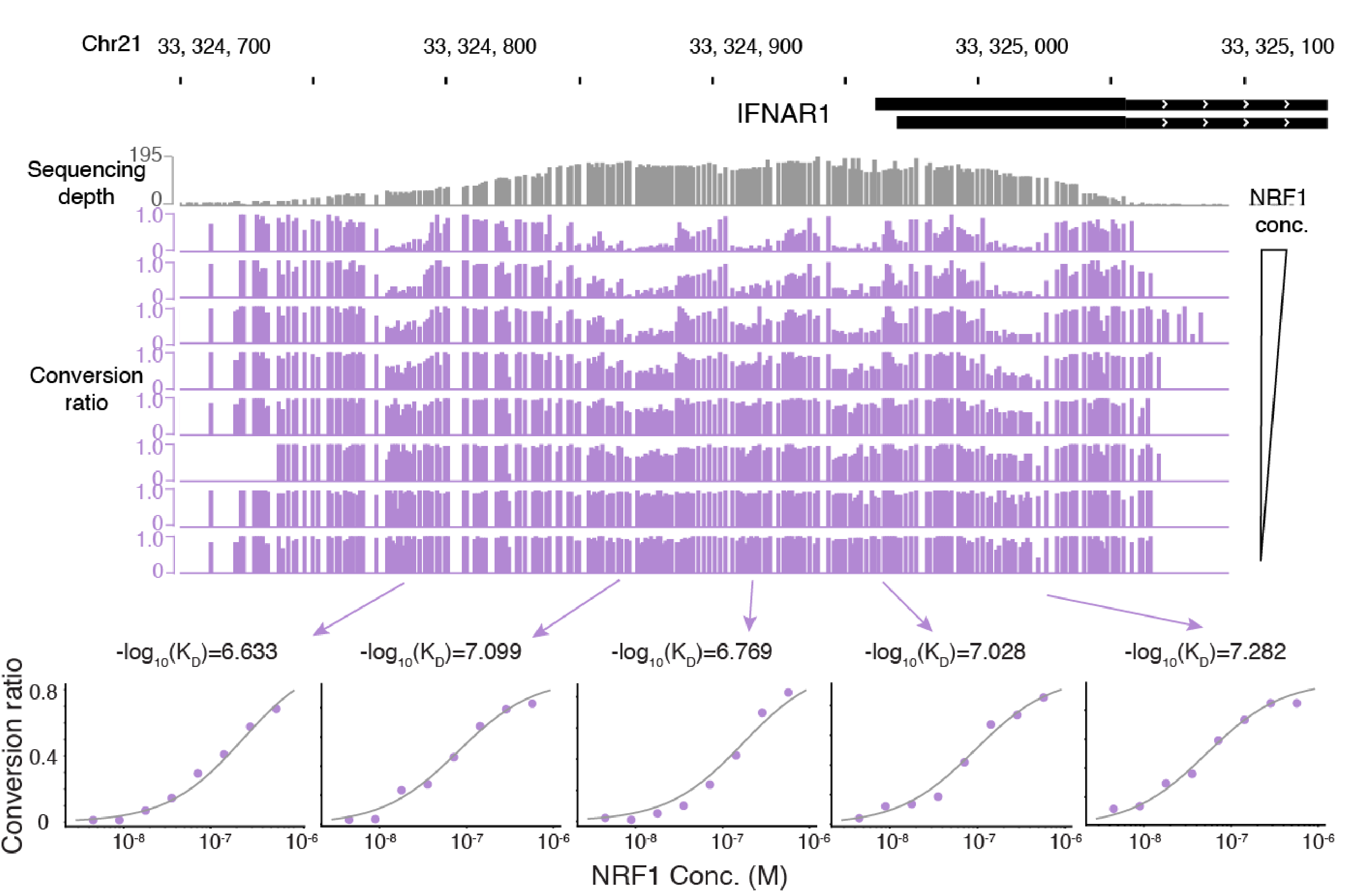
ivtFOODIE measures KDs of a purified transcriptional factor on its binding sites along the accessible genome
We recently extended FOODIE to an in vitro setting, ivtFOODIE, to directly measure equilibrium dissociation constants (KD) values across the accessible genome of a human cell line. For a given TF, we determine KD values at all binding sites by titrating 10 TF concentrations.
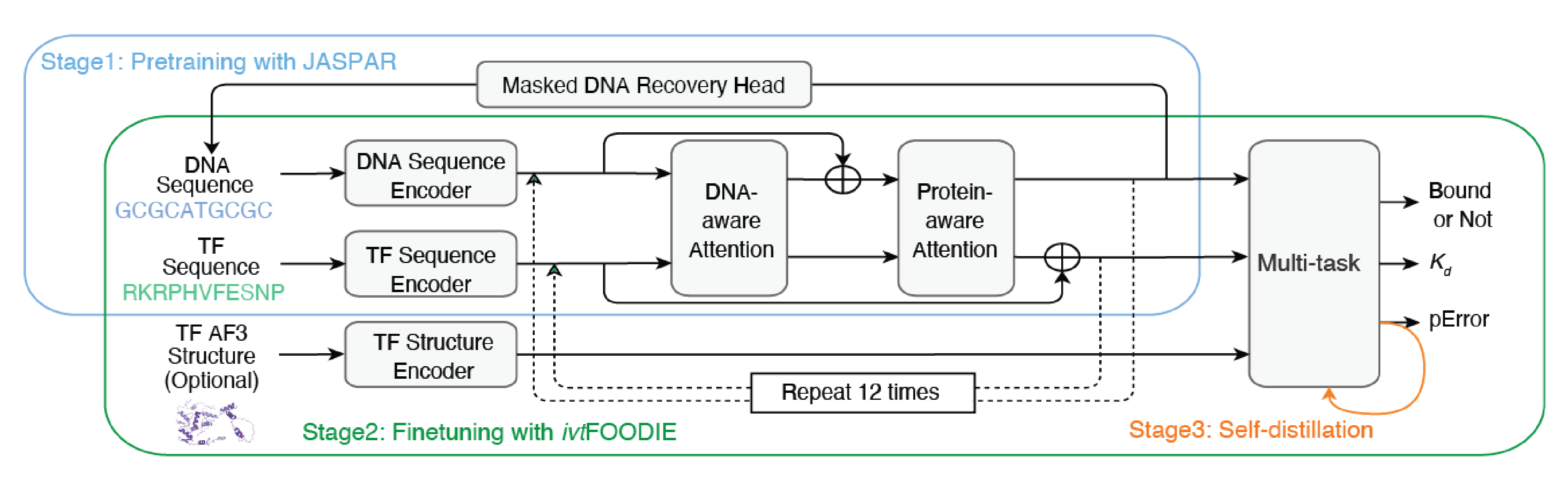
Prediction of any transcription factor’s binding affinity with any DNA sequence by deep learning with in vitro FOODIE data and public domain’s data
To reach accurate KD predictions of any TF on any DNA sites, we used the genome-wide KD data, together with the public domain binding motifs to successfully develop a deep learning model, Seq2 KD. Furthermore, we showed that mutations causal for certain diseases using Seq2 KD, such as cancer, often exhibit large KD changes on TF binding sites.
Single-cell FOODIE enables detection of cell-type-specific TF footprints in a pure cell population in a heterogeneous tissue, such as brain
By high-throughput single-cell FOODIE for cell typing, we have obtained transcription factor binding footprints across various cell types of human brain.
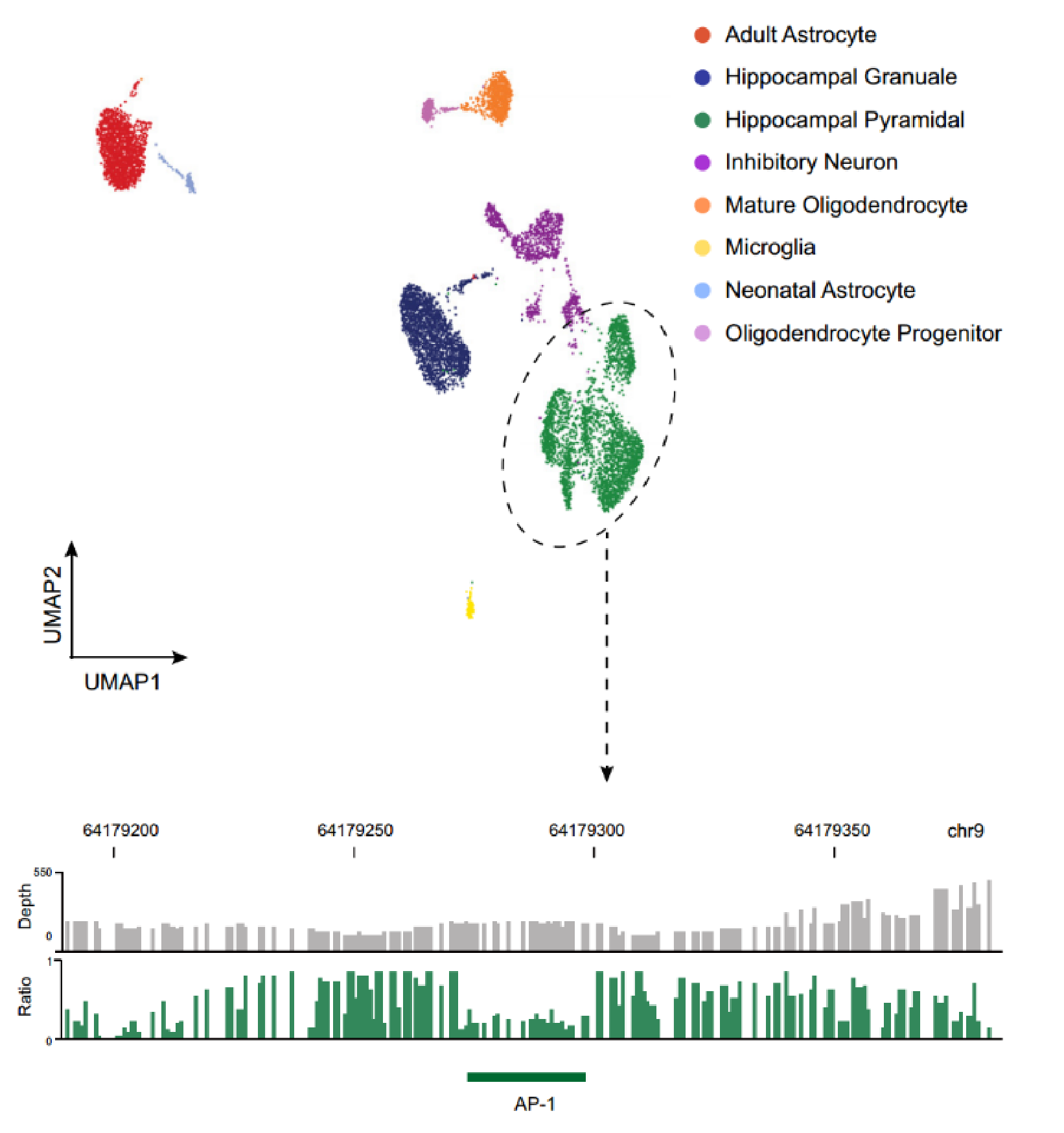
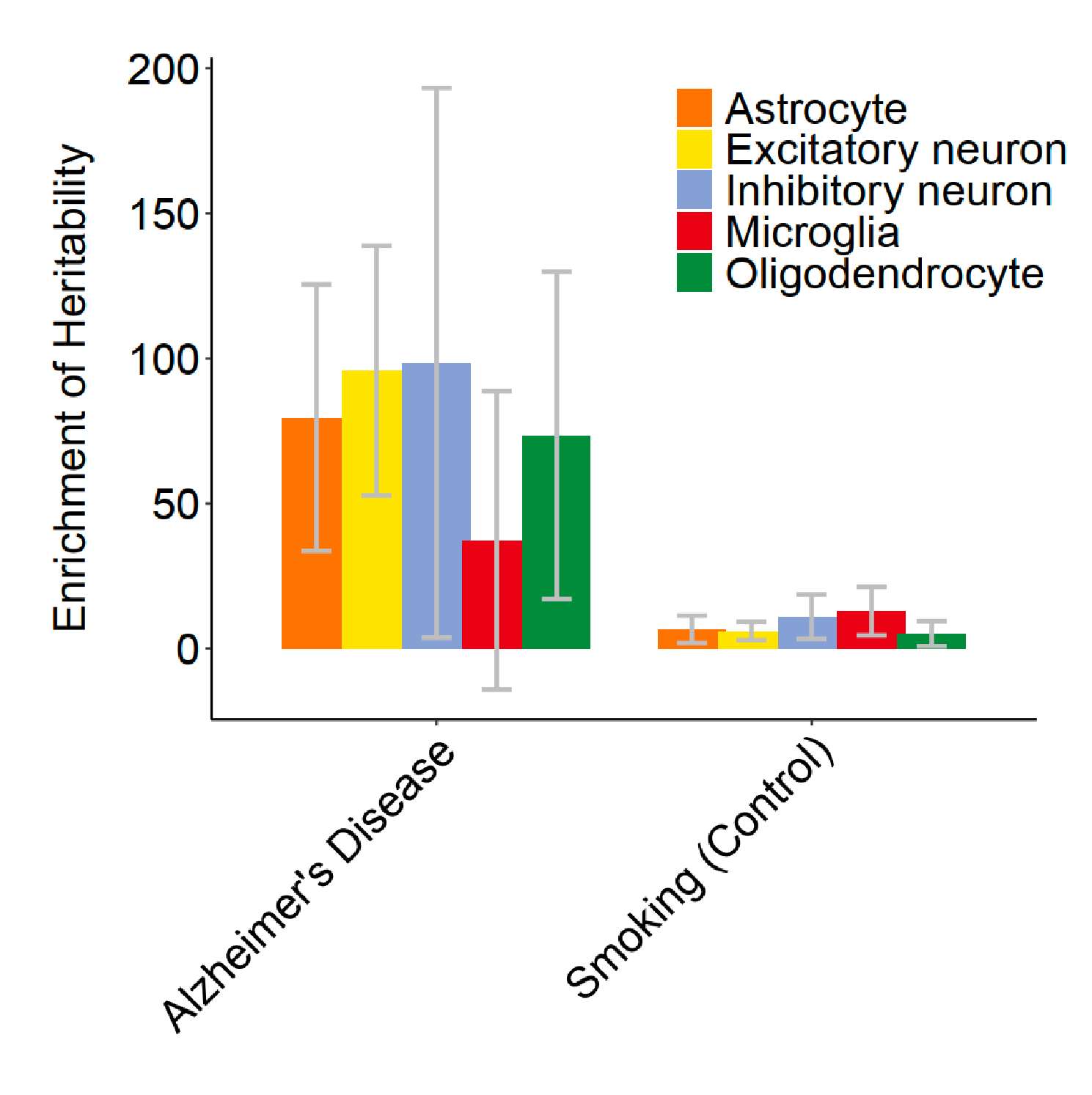
Alzheimer’s disease related risk variants are strongly enriched in the FOODIE TF footprints of the main cell types of human brain
We found GWAS SNVs associated with Alzheimer’s disease (AD) are highly enriched within TF footprints measured in the specific brain cell types. This finding highlights the use of FOODIE and disease cohort to link genotypic and phenotypic data for identifying potential drug targets for AD.