The canonical cytoskeleton is a three-dimensional network consisting of microtubule, long F-actin and intermediate filament. By contrast, the membrane skeleton is a spectrin-based two-dimensional network widely present in different metazoan cells. It is distributed underneath and parallel to the plasma membrane, endowing the membrane with high mechanical strength and plasticity (Figure 1). The membrane skeleton regulates the clustering of membrane proteins on the membrane and participate in the cellular response to extracellular signals. The membrane skeleton was first discovered in erythrocytes in the 1960s, and was later confirmed in various cell types including neurons, epithelial cells and lymphocytes. In neurons, it plays critical roles in the formation of axon initial segments and nodes of Ranvier, which are important for the initiation and rapid propagation of action potential.
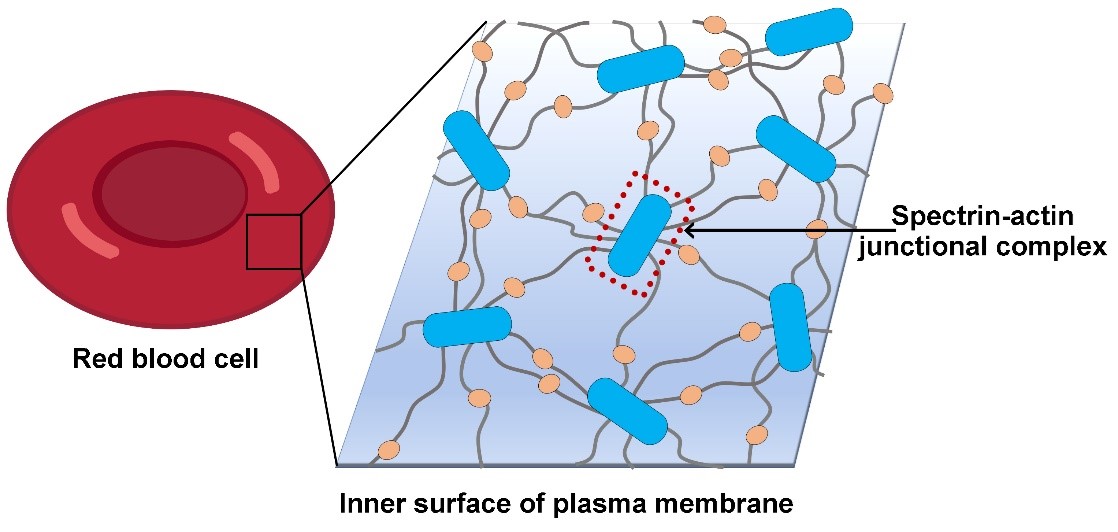
Figure 1, schematic diagram of erythrocyte membrane skeleton
In mammalian erythrocytes, the membrane skeleton is a hexagon-dominated polygonal 2D lattice, using a specific short F-actin as lattice nodes and a spectrin tetramer as lattice lines (Figure 1). At each node, F-actin acts as a hub to gather multiple spectrin fibers. A set of F-actin binding factors stabilize and regulate the activity of this spectrin-actin junction, including tropomyosin, tropomodulin, protein 4.1 (P4.1) and dematin. The lattice node is thereby named spectrin-actin junctional complex, which is the core complex of the skeleton. Up to date, the fine structure of the junctional complex is still lacking, which has limited the understanding of the organizational principle of the membrane skeleton and the exact roles of skeleton factors in membrane skeleton assembly and dynamics.
In a paper entitled “Structural basis of membrane skeleton organization in red blood cells” published on Cell, on April 11, 2023, a research team led by Prof. Ning Gao in Changping laboratory performed an ex vivo structural analysis of the membrane skeleton in its intact networked form (isolated from pig red blood cells) using cryo-EM, and determined the high-resolution structures of the spectrin-actin junctional complex.
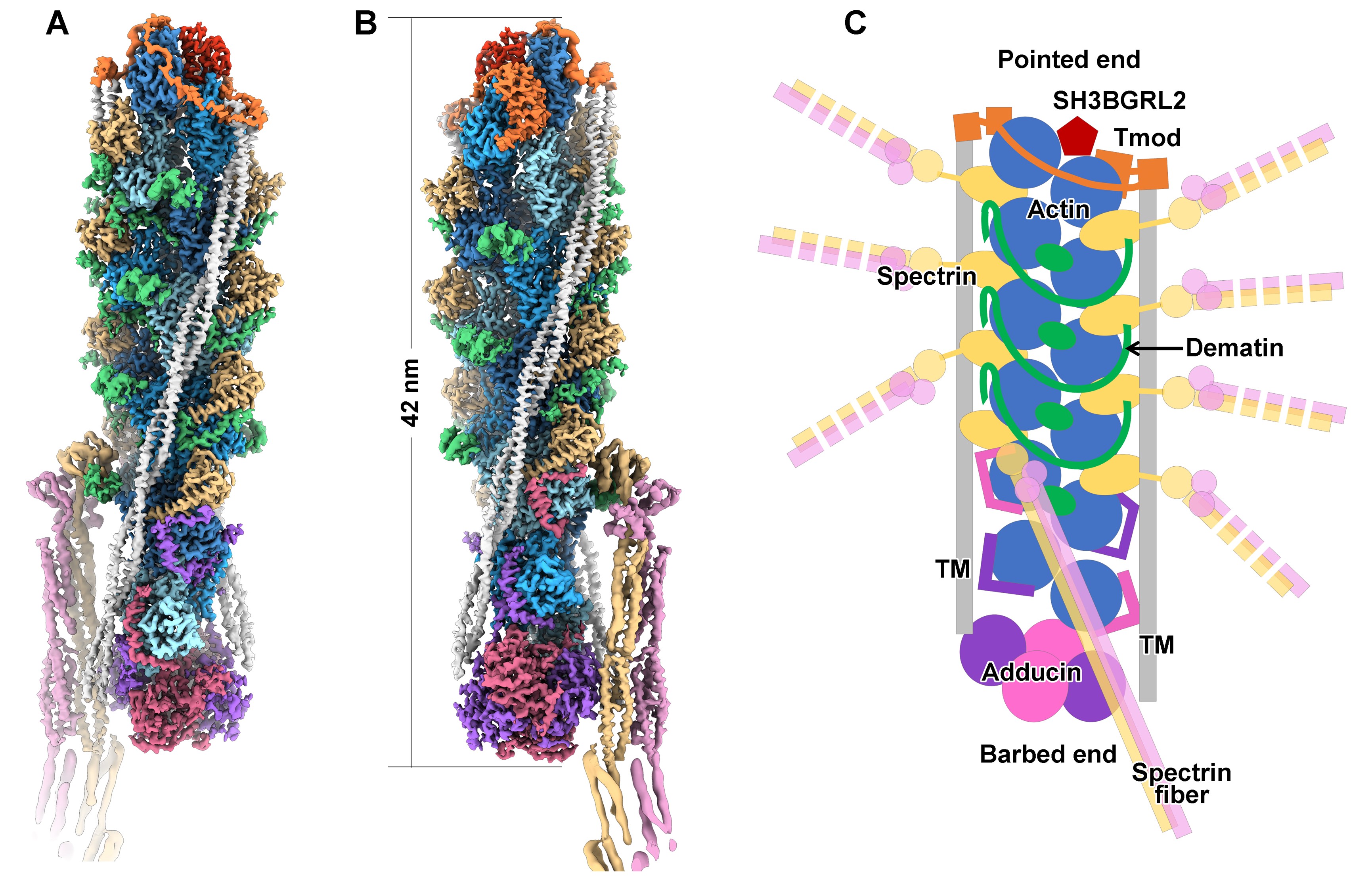
Figure 2, cryo-EM structure of the spectrin-actin junctional complex
The high-resolution structures demonstrate the organization details of the junctional complex (Figure 2). The core of the junctional complex is a six-layered short F-actin composed of 12 actin subunits, showing an overall length of ~42 nm. A total of 8 spectrin fibers were docked onto the F-actin at the inter-layer regions of layer 1 to layer 5 (from top to bottom in Figure 2). The two capping factors, tropomyosin and adducin, cap the two ends of F-actin (pointed end and barbed end), respectively, preventing the polymerization or depolymerization of F-actin. Two tropomyosin coiled coils are alongside the two strands of F-actin, and also interact with tropomodulin and adducin. The three factors together determine the length of the junctional complex. At the inter-layer regions of layer 2 to layer 5, three dematin molecules form ring-like structures encircling the trunk of the junctional complex to enhance the strength of the spectrin-actin junction. As a whole, these components of the junctional complex form an inter-connected network of inter-molecular interactions to stabilize the spectrin-actin junction.
In addition, this work presents several other novel findings: (1) The identification of SH3BGRL2 as a novel component of the junctional complex, being a capping protein of the pointed-end of the F-actin. (2) The identification of the tentacle-like actin-binding motifs in the flexible N- and C-terminal sequences of adducin proteins. (3) A novel docking mode of Tropomyosin on the F-actin, sharply different from the canonical one in the actomyosin system.
Overall, this work reveals the organization details of the junctional complex and membrane skeleton at molecular level, elucidate the molecular mechanism of skeleton factors in the membrane skeleton maintenance, and provide a structural framework to understand the assembly and dynamics of membrane skeleton.
This work was mainly conducted by Prof. Ning Gao, Dr Ningning Li and Ms Siyi Chen, and also contributed by Prof. Qiangfeng Cliff Zhang and Dr Kui Xu from Tsinghua University and Prof. Meng-Qiu Dong and Ms Meng-Ting He from National Institute of Biological Sciences, Beijing.